This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is transcriptomics?
Transcriptomics is the study of RNA molecules within the cell[1]. These RNA molecules are transcribed from the DNA template. There are many different types of RNA that can be studied but often mRNA (messenger RNA) is used, because it reflects the genes that are actively being expressed.
What techniques are used in transcriptomics?[1]
In general, the RNA must be first reverse-transcribed into cDNA (complimentary DNA). Reverse transcription is when RNA is transcribed into DNA. The cDNA is then used to analyze expression. Two common methods that are used to analyze expression levels are microarrays and RNA-seq.
Microarrays
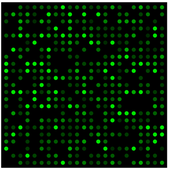
Microarrays have a smooth surface with DNA sequences of interest bound to it. cDNA with fluorescent markers is then washed over the surface of the slide. When this occurs, any cDNA that is complimentary to the DNA sequences of interest will bind. The level of fluorescence then indicates the level of expressions, because spots with higher amounts of binding will have brighter fluorescence.
Microarrays
Microarrays have a smooth surface with DNA sequences of interest bound to it. cDNA with fluorescent markers is then washed over the surface of the slide. When this occurs, any cDNA that is complimentary to the DNA sequences of interest will bind. The level of fluorescence then indicates the level of expressions, because spots with higher amounts of binding will have brighter fluorescence.
RNA-seq
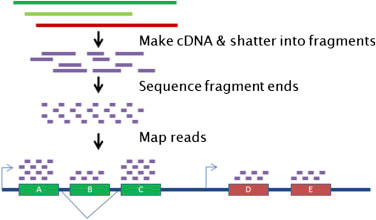
RNA-seq (RNA sequencing) is a newer technique that is more complex but also more flexible. In RNA-seq, RNA is first isolated from the tissue. It is then reverse transcribed into cDNA. The cDNA is then sequenced in short reads, and reassembled using bioinformatics tools. RNA-seq is convenient because it allows for analysis of the entire transcriptome (all RNA), rather than probing for specific targets.
RNA-seq (RNA sequencing) is a newer technique that is more complex but also more flexible. In RNA-seq, RNA is first isolated from the tissue. It is then reverse transcribed into cDNA. The cDNA is then sequenced in short reads, and reassembled using bioinformatics tools. RNA-seq is convenient because it allows for analysis of the entire transcriptome (all RNA), rather than probing for specific targets.
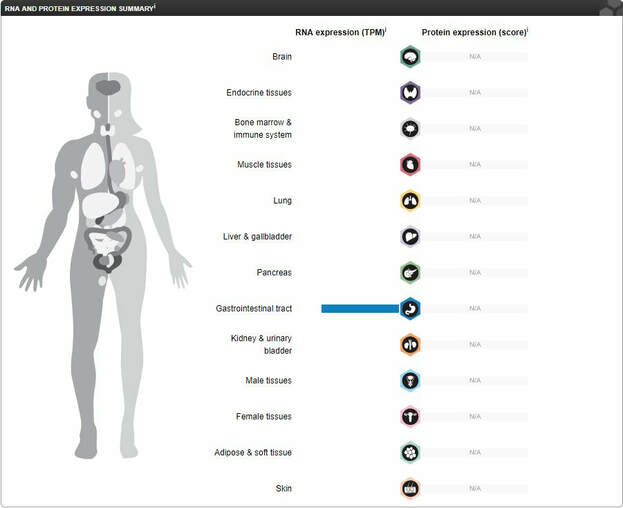
Where is LCT RNA expressed?
Conclusions
LCT, as it is specifically involved in polysaccharide digestion, has not been widely studied. The RNA expression profile has been quantified, and it is found only in the gastrointestinal tract, suggesting that it not involved in any other biological pathways other than digestion. However, there have not been significant studies on the upregulation or downregulation of other genes when LCT is mutated or knocked out, which could be interesting to learn about interactions with other genes and their respective gene ontologies.
References
[1] Blackburn, L. (2017, August). What is transcriptomics? University of Cambridge. www.phgfoundation.org/blog/what-is-transcriptomics
Header image: www.sciencemag.org/custom-publishing/webinars/power-rna-broad-application-rna-based-sequencing-transcriptome-and-genome
[1] Blackburn, L. (2017, August). What is transcriptomics? University of Cambridge. www.phgfoundation.org/blog/what-is-transcriptomics
Header image: www.sciencemag.org/custom-publishing/webinars/power-rna-broad-application-rna-based-sequencing-transcriptome-and-genome