This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are post translational modifications?
Post translational modifications are changes that are performed on the protein once it has been translated from mRNA into an amino acid chain[1]. The modifications are performed on specific residues (amino acids) within the protein. These modifications can significantly alter the structure and function of proteins, which results in a large increase in the size of the proteome.
What types of post translational modifications occur? [2]
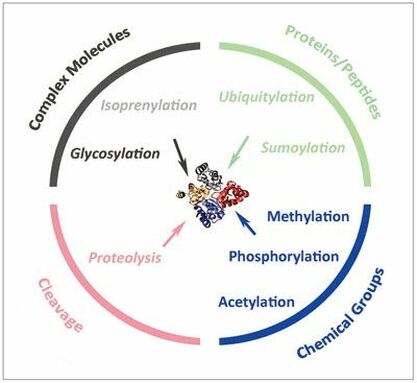
There are many different types of post translation modifications. Some are reversible, while others are permanent. Some of the most common types of post translational modifications are methylation, phosphyorylation, glycosylation, and acetylation.
Phyosphorylation is commonly performed on hydroxyl group of serine, tyrosine, and theronine. It also can occur on arginine, histidine, lysine, aspartate, and cysteine. Phosphorylation is a key mechanism in signaling.
Methylation commonly occurs on leucine, lysine, arginine, histidine, and glutamate. Methylation can modulate the activity of the target protein, and affect protein-protein interactions
Phyosphorylation is commonly performed on hydroxyl group of serine, tyrosine, and theronine. It also can occur on arginine, histidine, lysine, aspartate, and cysteine. Phosphorylation is a key mechanism in signaling.
Methylation commonly occurs on leucine, lysine, arginine, histidine, and glutamate. Methylation can modulate the activity of the target protein, and affect protein-protein interactions
Are there any post translational modifications of R1587 in LCT?
There are no predicted post translational modifications of arginine at position 1587 of LCT[3]. Although phosphorylation and methylation are both possible, neither are expected at a high threshold in LCT. Additionally, there is no expected phosphorylation or methylation expected at the substituted histidine at position 1587.
Conclusions
The single amino acid substitution R1587H in LCT is not expected to cause significant differences in post translational modifications. Niether the wildtype nor the mutant site is expected to be subject to the most common modifications: phosphorylation and methylation.
References
[1] Overview of Post-Translational Modifications (PTMs). Thermo Fisher Scientific. www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[2] Modified Residue. UniProt. www.uniprot.org/help/mod_res
[3] Artimo P, et al. (2012). ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res.
Header image: www.rcsb.org/structure/1OCC
[1] Overview of Post-Translational Modifications (PTMs). Thermo Fisher Scientific. www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
[2] Modified Residue. UniProt. www.uniprot.org/help/mod_res
[3] Artimo P, et al. (2012). ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res.
Header image: www.rcsb.org/structure/1OCC
